Sites description
Tiantishan Grottoes, known famously as “Liangzhou Grottoes” in the history, are located in the north of the Tiantishan, approximately 60 km south of the Wuwei City, which is a typical northwest city located in the eastern Hexi Corridor in Gansu Province, China (Fig. 7 A). The caves of the Tiantishan Grottoes were built in the Northern Liang (401-439AD) of sixteen kingdoms period about 1,600 years ago. It not only played an important role in spreading Buddhism and propagating Buddhist art in the Hexi Corridor region on the Silk Road, but also had profound influence on art characteristics of the grottoes in later generations, including the World Heritage Sites Yungang Grottoes and Longmen Grottoes, both of them are representative grottoes in China. Therefore, Tiantishan Grottoes was called the “Origin of Grottoes” by academic field.
Tiantishan Grottoes are inland and situated in the basin valley between the Qilian Mountains and the south mountains of Hexi corridor. It has a cold climate of semi-arid and high land with an average annual temperature of 4.9 ℃, annual rainfall of 159 mm. Because of the historical factors [53], most priceless historical objects including wall paintings and painted sculptures of the Tiantishan Grottoes were relocated to the Gansu Provincial Museum for the ex-situ protection in 1959, only a few mutilated remains of the painted sculptures or wall paintings were left in the original Tiantishan Grottoes. Began from 2006, these historical objects were returned gradually. However, due to seismic damage, festinate relocation, immature exfoliating technology, poorly conservation environment in the history, these priceless cultural objects suffered from various diseases, including paint losses, detachment, flaking, and microbial deterioration. In this case, the immediate and salvageable restoration is under way. Therefore, these objects which transferred to the Western Xia Museum of Wuwei city from Gansu Museum would then return to the Tiantishan Grottoes after completion of restoration.
Sampling process
Four sites were selected for the sampling of aerosol in four seasons (represented by April, June, October, and December) in 2016, including Cave 13 and Cave 18 of the Tiantishan Grottoes, warehouse inside and square outside of the Western Xia Museum, sample sites are namely TT13, TT18, TMI, and TMO, respectively (Fig. 7 B-D). In Cave 13, there was a giant sitting Shakyamuni Statue of 23.8 m which protected from the water by a semi-enveloping cofferdam constructed by reinforced concrete. Cave 18, only a central tower with a Buddha Statue remained its position is higher than Cave 13 with an open and flat landform. Cave 13 and cave 18 were two sites of the Tiantishan Grottoes which is still with surviving wall paintings. Western Xia Museum is located in the southeast of the Confucius' temple in Wuwei City, the square outside of it and the surrounding areas are always with activities of people and vehicles, however, the warehouse inside of the museum has been employed to store the uncovered wall paintings of the Tiantishan Grottoes, in where these artworks have been gradually restored in recent years. It should be noted that the restoring work of the cultural relics was only carried out during the period from April to November in a year and suspended at rest for the months without human disturbance due to the local bitter-cold weather.
Buck Bio-Culture™ Model B30120 sampler (A.P. Buck Inc, USA) was used to sampling from the atmosphere in Maijishan Grottoes. At each site, the sampler was installed 1.5 m above ground level with a supporting platform. Samples were carried out for 2 min, with three parallel repetitions. For each sampling, the bio-aerosol sampler loaded with 90 mm Petri dishes containing Potato Dextrose Agar (PDA) medium. Exposed culture dishes were incubated for 7-15 days at room temperature with 25 ℃.
Fungal identification
Colony forming units (CFU) on the plates were enumerated, and fungal concentrations were expressed as per cubic meter of air (CFU/m3). CFU/m3 was calculated as below:
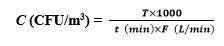
In the equations,C is the airborne fungi concentration; T is the total colonies on the PDA plates; t is the sampling time; F is the airflow rate.
After incubating, microbial colonies were counted and fungi were preliminary identified according to morphological characteristics, including shape, size, color, and refractivity, etc. The isolates were identified by using a molecular method as described below. Each isolate was homogenized in liquid nitrogen and then genomic DNA was extracted by using a commercially available extraction Kit (Tiangen Co., Beijing, China) according to the manufacturer's protocols. The internal transcribed spacer (ITS) region of fungal rRNA genes was amplified with the following universal primer set: ITS1 (5'-TCCGTAGGTGAACCTGCGG-3') and ITS4 (5'-TCCTCCGCTTATTGATATGC-3') [54]. The reaction mixture (25 μL) consisted of 2.5 μL 10 × PCR buffer, 1-unit Taq polymerase (Tiangen Co., Beijing, China), 0.2 mM dNTPs, 2.5 mM MgCl2, 0.2 μM each primer, and 2.5 μL (ca. 10 ng DNA) template. The PCR program included an initial denaturation at 94 ℃ for 5 min, 30 cycles at 94 ℃ for 40 s, annealing at 55 ℃ for 40 s, an extension at 72 ℃ for 40 s, and a final extension step at 72 ℃ for 10 min. The PCR products were detected by electrophoresis on 1.0% agarose gel.
The similarities among PCR products were determined by restriction fragment length polymorphisms (RFLP) analysis. The PCR products were digested with the double restriction endonucleases BsuR I and Hinf I (MBI, Fermentas). And then, the digested fragments were differentiated into several clusters according to their spectral patterns on 2.5% agarose gel.
Cloning of PCR product of distinctive strain was performed with the pGEM-T Vector System (Tiangen Co., Beijing, China) following their manufacturer’s instructions, and the ligation product was subsequently transformed in Escherichia coli DH5α competent cells (Tiangen Co., Beijing, China), which allows Blue-White selection. The transformants were plated on LB medium with ampicillin (100 mg/ml), X-Gal (20 mg/ml) and IPTG (200 mg/ml). Positive clones were identified by PCR amplification with the pGEM-T vector primer pairs (T7/SP6) by using the same program as ITS amplification.
Fungal suspensions of expected clones were sequenced by the Shanghai Majorbio Bio-technology Co., Ltd. (Shanghai, China). A total of 34 fungi representative sequences were obtained (ca. 600 bp) and then analyzed via National Center for Biotechnology Information (NCBI) Blast program (https://blast.ncbi.nlm.nih.gov/Blast.cgi). The most similar sequences were extracted from the GenBank database. A phylogenetic neighbor-joining tree was constructed via the MEGA software 7.0. The sequences retrieved during this study could be accessed under the data range of MH042805-MH042838.
Data of environmental parameters
The meteorological data were collected in all four sampling sites for one year (from April 16, 2016 to April 16, 2017). Temperature (T, °C) and relative humidity (RH, %) were collected by HOBO® U23-001 Temp/RH data logger (Onset Computer Corporation, USA). Data of rainfall was provided by local weather bureaus. For all environmental parameters, the values used in data analyses were 10-day averages from before and after the sampling days.
Statistical analysis
All the experiments were analyzed by one-way analysis of variance (ANOVA) via SPSS version 16.0. The relationships between airborne fungi concentration and environmental parameters were tested via Pearson correlation analysis. Shannon-Weiner diversity index of fungal communities were generated using the Vegan packages in R (4.0.0). Canonical correlation analysis (CCA) for airborne fungal communities and environmental parameters were then analyzed using CANOCO version 4.5.